Fit growth models and extract growth statistics#
This tutorial demonstrates how to fit growth models and extract growth statistics using the growthcurves package.
The analysis workflow includes:
Generating or loading growth data
Fitting mechanistic models (ODE-based, parametric)
Fitting phenomenological models (parametric and non-parametric)
Extracting growth statistics from all fits
Saving results for visualization
For preprocessing examples (blank subtraction, outlier detection, path length correction), see the companion notebook:
preprocessing.ipynb.
For visualization of the results, see the companion notebook:
plotting.ipynb (Visualize fitted growth curves, derivatives,
and growth statistics)
from pprint import pprint
import numpy as np
import pandas as pd
import growthcurves as gc
Generate synthetic data#
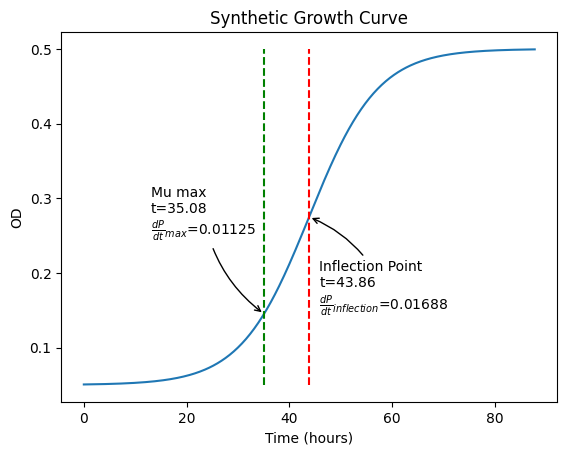
This cell generates synthetic growth data from a clean logistic function.
time is modeled in hours, with measurements every 12 minutes (0.2 hours) for a total of 440 points (88 hours).
We assume a lag of 30 hours, an intrinsic growth rate of 0.15 hour⁻¹, and a carrying capacity of 0.45 OD.
Doubling time at maximum acceleration point (t=35.08): 5.86 hours
Maximum specific growth rate (mu_max of log curve) at t=35.08: 0.11830 hour^-1
How Growth Parameters Are Calculated#
The table below summarizes how the main reported growth statistics are calculated across model classes.
Output key |
Meaning |
How it is calculated |
|---|---|---|
|
Maximum observed/fitted OD |
Maximum OD over the valid data range |
|
Maximum specific growth rate (μ_max) |
Maximum of |
|
Intrinsic model rate parameter |
For mechanistic models: fitted intrinsic |
|
Doubling time in hours |
|
|
Time at maximum specific growth |
Time where |
|
OD at time of μ_max |
Model-predicted OD at |
|
Exponential phase boundaries |
From threshold or tangent phase-boundary method in |
|
Fit error |
RMSE between observed OD and model-predicted OD over the model fit window |
For this tutorial:
Mechanistic comparisons use mechanistic parametric fits.
Phenomenological comparisons include both phenomenological parametric and non-parametric fits.
Extract growth stats from the dataset#
The extract_stats_from_fit() function calculates these key metrics:
max_od: Maximum OD value within the fitted windowmu_max: Observed maximum specific growth rate μ_max (hour⁻¹) - calculated from the fitted curveintrinsic_growth_rate: Model parameter for intrinsic growth rate (parametric models only,Nonefor non-parametric)doubling_time: Time to double the population at peak growth (hours)exp_phase_start: When exponential phase begins (hours)exp_phase_end: When exponential phase ends (hours)time_at_umax: Time when μ reaches its maximum (hours)od_at_umax: OD value at time of maximum μfit_t_min: Start of fitting window (hours)fit_t_max: End of fitting window (hours)fit_method: Identifier for the method usedmodel_rmse: Root mean squared error
Descriptive parameters are extracted from the fits. Where parameters are not extracted directly from the fitted model, they are calculated. The table below shows how different stats are calculated according to the different approaches:
MECHANISTIC MODELS#
Name |
Model |
Equation |
Exp Start |
Exp End |
Intrinsic μ |
μ max |
Carrying Capacity |
Fit |
|---|---|---|---|---|---|---|---|---|
Logistic |
parametric |
|
threshold/ |
threshold/ |
μ |
max dln(N)/dt |
K |
entire curve |
Gompertz |
parametric |
|
threshold/ |
threshold/ |
μ |
max dln(N)/dt |
K |
entire curve |
Richards |
parametric |
|
threshold/ |
threshold/ |
μ |
max dln(N)/dt |
A |
entire curve |
Baranyi |
parametric |
|
threshold/ |
threshold/ |
μ |
max dln(N)/dt |
K |
entire curve |
PHENOMENOLOGICAL MODELS#
Name |
Model |
Equation |
Exp Start |
Exp End |
Intrinsic μ |
μ max |
Max OD |
Fit |
|---|---|---|---|---|---|---|---|---|
Linear |
non-parametric |
|
threshold/ |
threshold/ |
n.a. |
b |
max OD raw |
only window |
Spline |
non-parametric |
|
threshold/ |
threshold/ |
n.a. |
max of derivative of spline |
max OD raw |
only log phase |
Logistic (phenom) |
parametric |
|
λ |
threshold/ |
n.a. |
μ_max |
K |
entire curve |
Gompertz (phenom) |
parametric |
|
λ |
threshold/ |
n.a. |
μ_max |
K |
entire curve |
Gompertz (modified) |
parametric |
|
λ |
threshold/ |
n.a. |
μ_max |
K |
entire curve |
Richards (phenom) |
parametric |
|
λ |
threshold/ |
n.a. |
μ_max |
K |
entire curve |
Understanding Growth Rates: Intrinsic vs. Observed#
Important distinction:
mu_max(μ_max): The observed maximum specific growth rate calculated from the fitted curve as max(d(ln N)/dt). This is what you measure from the data.intrinsic_growth_rate: The model parameter representing intrinsic growth capacity:Parametric models: This is a fitted parameter (e.g.,
rin Logistic,mu_maxin Gompertz)Non-parametric methods: Returns
None(no model parameter exists)
Mechanistic Models#
Mechanistic models are ODE-based parametric models that encode growth dynamics as differential equations.
Fit Models#
# Fit mechanistic models
fit_mech_logistic = gc.parametric.fit_parametric(t, N, method="mech_logistic")
fit_mech_gompertz = gc.parametric.fit_parametric(t, N, method="mech_gompertz")
fit_mech_richards = gc.parametric.fit_parametric(t, N, method="mech_richards")
fit_mech_baranyi = gc.parametric.fit_parametric(t, N, method="mech_baranyi")
# Combine fits into a dictionary
mechanistic_fits = {
"mech_logistic": fit_mech_logistic,
"mech_gompertz": fit_mech_gompertz,
"mech_richards": fit_mech_richards,
"mech_baranyi": fit_mech_baranyi,
}
# Display example fit result
print("=== Logistic Fit Result ===")
pprint(fit_mech_logistic, indent=2)
=== Logistic Fit Result ===
{ 'model_type': 'mech_logistic',
'params': { 'K': np.float64(0.4499768424051535),
'N0': np.float64(0.0006250371636641061),
'fit_t_max': 87.8,
'fit_t_min': 0.0,
'mu': np.float64(0.1499259523625051),
'y0': np.float64(0.050009668376952275)}}
Extract Growth Statistics#
# Extract stats from each mechanistic fit
stats_mech_logistic = gc.inference.extract_stats(fit_mech_logistic, t, N)
stats_mech_gompertz = gc.inference.extract_stats(fit_mech_gompertz, t, N)
stats_mech_richards = gc.inference.extract_stats(fit_mech_richards, t, N)
stats_mech_baranyi = gc.inference.extract_stats(fit_mech_baranyi, t, N)
# Combine stats into a dictionary
mechanistic_stats = {
"mech_logistic": stats_mech_logistic,
"mech_gompertz": stats_mech_gompertz,
"mech_richards": stats_mech_richards,
"mech_baranyi": stats_mech_baranyi,
}
# Display growth statistics for logistic fit
print("=== Logistic Growth Statistics ===")
pprint(stats_mech_logistic, indent=2)
# Create comparison dataframe
print("\n=== Mechanistic Models Comparison ===")
mechanistic_df = pd.DataFrame(mechanistic_stats).T[
[
"mu_max",
"intrinsic_growth_rate",
"doubling_time",
"time_at_umax",
"exp_phase_start",
"exp_phase_end",
"model_rmse",
]
]
mechanistic_df.T
=== Logistic Growth Statistics ===
{ 'N0': 0.050009668376952275,
'doubling_time': 8.861181001951804,
'exp_phase_end': 59.48715239678405,
'exp_phase_start': 12.915015487973315,
'fit_method': 'model_fitting_mech_logistic',
'fit_t_max': 87.8,
'fit_t_min': 0.0,
'intrinsic_growth_rate': 0.1499259523625051,
'max_od': 0.49998651078210576,
'model_rmse': 4.597890449408753e-05,
'mu_max': 0.0782228892974051,
'od_at_umax': 0.15665653300529114,
'time_at_umax': 36.07014028056112}
=== Mechanistic Models Comparison ===
| mech_logistic | mech_gompertz | mech_richards | mech_baranyi | |
|---|---|---|---|---|
| mu_max | 0.078223 | 0.081681 | 0.081759 | 0.077833 |
| intrinsic_growth_rate | 0.149926 | 0.055765 | 0.187789 | 0.149806 |
| doubling_time | 8.861181 | 8.486063 | 8.477922 | 8.905564 |
| time_at_umax | 36.07014 | 22.873747 | 35.542285 | 36.246092 |
| exp_phase_start | 12.915015 | 6.520981 | 15.613263 | 1.655978 |
| exp_phase_end | 59.487152 | 65.012076 | 59.727944 | 59.522783 |
| model_rmse | 0.000046 | 0.02491 | 0.001558 | 0.00004 |
Phenomenological Models - Parametric#
These are phenomenological parametric models fit in ln-space.
Fit Models#
# Fit phenomenological parametric models
fit_phenom_logistic = gc.parametric.fit_parametric(t, N, method="phenom_logistic")
fit_phenom_gompertz = gc.parametric.fit_parametric(t, N, method="phenom_gompertz")
fit_phenom_gompertz_modified = gc.parametric.fit_parametric(
t, N, method="phenom_gompertz_modified"
)
fit_phenom_richards = gc.parametric.fit_parametric(t, N, method="phenom_richards")
# Combine fits into a dictionary
phenom_param_fits = {
"phenom_logistic": fit_phenom_logistic,
"phenom_gompertz": fit_phenom_gompertz,
"phenom_gompertz_modified": fit_phenom_gompertz_modified,
"phenom_richards": fit_phenom_richards,
}
# Display example fit
print("=== Phenomenological Logistic Fit ===")
pprint(fit_phenom_logistic, indent=2)
=== Phenomenological Logistic Fit ===
{ 'model_type': 'phenom_logistic',
'params': { 'A': np.float64(2.299980001744068),
'N0': np.float64(0.050273088569627304),
'fit_t_max': 87.8,
'fit_t_min': 0.0,
'lam': np.float64(21.923803883807267),
'mu_max': np.float64(0.07997940768473631)}}
Extract Growth Statistics#
# Extract stats from each phenomenological parametric fit
stats_phenom_logistic = gc.inference.extract_stats(
fit_phenom_logistic, t, N, phase_boundary_method="tangent"
)
stats_phenom_gompertz = gc.inference.extract_stats(
fit_phenom_gompertz, t, N, phase_boundary_method="tangent"
)
stats_phenom_gompertz_modified = gc.inference.extract_stats(
fit_phenom_gompertz_modified, t, N, phase_boundary_method="tangent"
)
stats_phenom_richards = gc.inference.extract_stats(
fit_phenom_richards, t, N, phase_boundary_method="tangent"
)
# Combine stats into a dictionary
phenom_param_stats = {
"phenom_logistic": stats_phenom_logistic,
"phenom_gompertz": stats_phenom_gompertz,
"phenom_gompertz_modified": stats_phenom_gompertz_modified,
"phenom_richards": stats_phenom_richards,
}
# Display example stats
print("=== Phenomenological Logistic Stats ===")
pprint(stats_phenom_logistic, indent=2)
# Create comparison dataframe
print("\n=== Phenomenological Parametric Models Comparison ===")
phenom_param_df = pd.DataFrame(phenom_param_stats).T[
[
"mu_max",
"intrinsic_growth_rate",
"doubling_time",
"time_at_umax",
"exp_phase_start",
"exp_phase_end",
"model_rmse",
]
]
phenom_param_df.T
=== Phenomenological Logistic Stats ===
{ 'N0': 0.050273088569627304,
'doubling_time': 8.666570566416299,
'exp_phase_end': 50.65869644960617,
'exp_phase_start': 21.923803883807267,
'fit_method': 'model_fitting_phenom_logistic',
'fit_t_max': 87.8,
'fit_t_min': 0.0,
'intrinsic_growth_rate': None,
'max_od': 0.5005310454634615,
'model_rmse': 0.0008165377630810733,
'mu_max': 0.07997940768473631,
'od_at_umax': 0.15805737079248627,
'time_at_umax': 36.246092184368734}
=== Phenomenological Parametric Models Comparison ===
| phenom_logistic | phenom_gompertz | phenom_gompertz_modified | phenom_richards | |
|---|---|---|---|---|
| mu_max | 0.079979 | 0.092212 | 0.089873 | 0.078659 |
| intrinsic_growth_rate | None | None | None | None |
| doubling_time | 8.666571 | 7.516896 | 7.71256 | 8.812088 |
| time_at_umax | 36.246092 | 33.782766 | 33.430862 | 36.597996 |
| exp_phase_start | 21.923804 | 25.07681 | 23.467609 | 21.241111 |
| exp_phase_end | 50.658696 | 48.666283 | 48.628298 | 50.891908 |
| model_rmse | 0.000817 | 0.005205 | 0.00408 | 0.000506 |
Phenomenological Models - Non-Parametric#
These are phenomenological non-parametric fits that estimate growth features directly from local trends and smoothing.
For spline fitting, use smooth to choose smoothing behavior:
smooth="fast"(default): auto-default lambda rulesmooth="slow": weighted GCV smoothingsmooth=<float>: manual lambda value
Fit Models#
# Fit non-parametric models
# Spline supports smooth="fast" (default), smooth="slow", or a manual float.
fit_spline = gc.non_parametric.fit_non_parametric(
t,
N,
method="spline",
smooth="fast",
)
# Example manual smoothing value:
# fit_manual = gc.non_parametric.fit_non_parametric(t, N, method="spline", smooth=0.5)
fit_sliding_window = gc.non_parametric.fit_non_parametric(
t,
N,
method="sliding_window",
window_points=7,
)
# Combine fits into a dictionary
phenom_nonparam_fits = {
"spline": fit_spline,
"sliding_window": fit_sliding_window,
}
# Display non-parametric fit results
pprint(phenom_nonparam_fits, indent=2)
{ 'sliding_window': { 'model_type': 'sliding_window',
'params': { 'fit_t_max': 36.8,
'fit_t_min': 35.6,
'intercept': -4.663248124939173,
'slope': 0.07789418101795426,
'time_at_umax': 36.2,
'window_points': 7}},
'spline': { 'model_type': 'spline',
'params': { 'fit_t_max': 87.8,
'fit_t_min': 0.0,
'mu_max': 0.07792405326815431,
'smooth': 'fast',
'spline_s': 1.061160014205202e-06,
'tck_c': [ -2.983329231043416,
-2.9832050748046637,
-2.98295676232716,
-2.9825702163662196,
-2.98217283102193,
-2.9817633571643003,
-2.9813416757982893,
-2.9809073679458216,
-2.980460078264196,
-2.9799994221830715,
-2.9795250092010335,
-2.979036436615012,
-2.9785332907673787,
-2.9780151463732953,
-2.977481566330003,
-2.9769321013969012,
-2.976366289901965,
-2.9757836574350187,
-2.9751837165379413,
-2.9745659663893727,
-2.9739298924846227,
-2.9732749663107403,
-2.9726006450169047,
-2.971906371080261,
-2.971191571967351,
-2.97045565979134,
-2.969698030965196,
-2.968918065851066,
-2.968115128406077,
-2.9672885658248327,
-2.966437708178866,
-2.965561868053396,
-2.9646603401817124,
-2.963732401077555,
-2.9627773086659075,
-2.9617943019126054,
-2.960782600453268,
-2.9597414042220236,
-2.9586698930805904,
-2.9575672264482686,
-2.956432542933481,
-2.9552649599675123,
-2.954063573441161,
-2.952827457345031,
-2.951555663414292,
-2.950247220778713,
-2.9489011356189017,
-2.947516390829657,
-2.946091945691482,
-2.944626735551251,
-2.9431196715132004,
-2.941569640141362,
-2.939975503174696,
-2.9383360972562182,
-2.936650233677407,
-2.9349166981393924,
-2.93313425053232,
-2.9313016247344845,
-2.92941752843277,
-2.927480642966142,
-2.9254896231938394,
-2.923443097390115,
-2.921339667167345,
-2.91917790742946,
-2.916956366357631,
-2.9146735654302875,
-2.9123279994795555,
-2.909918136786242,
-2.907442419215605,
-2.904899262396145,
-2.9022870559436926,
-2.899604163733205,
-2.8968489242204876,
-2.89401965081653,
-2.8911146323166195,
-2.8881321333866703,
-2.8850703951094916,
-2.881927635593229,
-2.878702050644505,
-2.875391814508704,
-2.8719950806798105,
-2.868509982782168,
-2.864934635526531,
-2.86126713574262,
-2.8575055634904425,
-2.8536479832524426,
-2.8496924452085453,
-2.84563698659592,
-2.8414796331553345,
-2.8372184006656047,
-2.8328512965677,
-2.828376321679734,
-2.8237914720039208,
-2.819094740626403,
-2.814284119710559,
-2.809357602584179,
-2.8043131859206754,
-2.7991488720141104,
-2.793862671147633,
-2.7884526040544984,
-2.7829167044706202,
-2.777253021777138,
-2.771459623731216,
-2.765534599282865,
-2.7594760614751888,
-2.7532821504251093,
-2.7469510363811223,
-2.7404809228543257,
-2.7338700498184596,
-2.7271166969743246,
-2.7202191870734858,
-2.713175889295768,
-2.7059852226746,
-2.6986456595639043,
-2.691155729139747,
-2.68351402092962,
-2.6757191883618594,
-2.667769952327276,
-2.659665104744799,
-2.6514035121225894,
-2.642984119105768,
-2.6344059520016567,
-2.625668122273214,
-2.616769829991103,
-2.6077103672347044,
-2.5984891214322445,
-2.5891055786301242,
-2.5795593266814945,
-2.569850058344123,
-2.5599775742776587,
-2.5499417859304256,
-2.539742718306133,
-2.5293805126009348,
-2.5188554287016087,
-2.50816784753583,
-2.4973182732658663,
-2.486307335317367,
-2.475135790235316,
-2.463804523359694,
-2.452314550313793,
-2.440667018298731,
-2.428863207188198,
-2.416904530418032,
-2.4047925356658535,
-2.3925289053165564,
-2.380115456710125,
-2.3675541421688497,
-2.3548470488016964,
-2.3419963980842184,
-2.3290045452130728,
-2.3158739782348374,
-2.3026073169494943,
-2.2892073115895384,
-2.2756768412763586,
-2.262018912256053,
-2.2482366559174998,
-2.2343333265960235,
-2.22031229916652,
-2.2061770664304476,
-2.19193123630153,
-2.1775785287954723,
-2.1631227728294036,
-2.1485679028371614,
-2.1339179552068184,
-2.119177064547257,
-2.1043494597907757,
-2.0894394601390873,
-2.0744514708601294,
-2.059389978943485,
-2.044259548622206,
-2.0290648167691008,
-2.0138104881755763,
-1.9985013307212784,
-1.9831421704428023,
-1.9677378865098611,
-1.9522934061172825,
-1.9368136993012832,
-1.9213037736884622,
-1.905768669186002,
-1.8902134526215275,
-1.8746432123411134,
-1.8590630527739378,
-1.8434780889720017,
-1.827893441133471,
-1.8123142291180303,
-1.7967455669627896,
-1.7811925574071625,
-1.7656602864352298,
-1.7501538178440303,
-1.7346781878462878,
-1.7192383997160126,
-1.7038394184855101,
-1.6884861657022106,
-1.6731835142538345,
-1.657936283270323,
-1.642749233110963,
-1.627627060445113,
-1.612574393434866,
-1.5975957870279422,
-1.5826957183690293,
-1.5678785823376644,
-1.5531486872206728,
-1.5385102505269874,
-1.5239673949525594,
-1.5095241445028122,
-1.4951844207799363,
-1.4809520394419953,
-1.4668307068405897,
-1.4528240168434547,
-1.4389354478480694,
-1.4251683599919107,
-1.4115259925646337,
-1.3980114616269468,
-1.3846277578405433,
-1.371377744512857,
-1.3582641558599768,
-1.345289595490387,
-1.3324565351116984,
-1.3197673134618961,
-1.3072241354660006,
-1.2948290716184256,
-1.2825840575906582,
-1.270490894063221,
-1.2585512467802507,
-1.2467666468243392,
-1.2351384911086667,
-1.2236680430827667,
-1.2123564336477082,
-1.201204662275782,
-1.190213598329267,
-1.1793839825722359,
-1.1687164288688405,
-1.1582114260610143,
-1.1478693400180449,
-1.1376904158500503,
-1.1276747802769833,
-1.1178224441444349,
-1.1081333050772078,
-1.0986071502613497,
-1.089243659345125,
-1.0800424074492094,
-1.071002868276299,
-1.0621244173101814,
-1.0534063350943457,
-1.044847810580157,
-1.0364479445346964,
-1.0282057529984592,
-1.0201201707832224,
-1.012190055000567,
-1.0044141886117433,
-0.9967912839898142,
-0.989319986485258,
-0.9819988779865393,
-0.9748264804674494,
-0.967801259513384,
-0.9609216278190884,
-0.9541859486507622,
-0.9475925392658469,
-0.9411396742841891,
-0.9348255890047206,
-0.9286484826621945,
-0.922606521618957,
-0.9166978424871488,
-0.9109205551771596,
-0.9052727458685921,
-0.8997524799003643,
-0.8943578045770575,
-0.8890867518889294,
-0.8839373411434722,
-0.8789075815067319,
-0.8739954744529654,
-0.8691990161215787,
-0.8645161995806084,
-0.8599450169963107,
-0.8554834617087599,
-0.8511295302135871,
-0.8468812240503051,
-0.8427365515978724,
-0.8386935297784015,
-0.8347501856701177,
-0.8309045580308804,
-0.8271546987337323,
-0.8234986741161345,
-0.8199345662446573,
-0.8164604740970577,
-0.8130745146637534,
-0.8097748239708298,
-0.8065595580267887,
-0.8034268936953078,
-0.8003750294963714,
-0.7974021863381299,
-0.7945066081819346,
-0.7916865626429664,
-0.7889403415289372,
-0.7862662613193072,
-0.7836626635874961,
-0.7811279153685148,
-0.7786604094744619,
-0.776258564760279,
-0.7739208263421362,
-0.7716456657707904,
-0.7694315811621912,
-0.7672770972876058,
-0.7651807656254356,
-0.7631411643768858,
-0.7611568984475742,
-0.7592265993971004,
-0.7573489253585592,
-0.7555225609299046,
-0.7537462170390014,
-0.752018630784171,
-0.7503385652519243,
-0.7487048093135675,
-0.7471161774022385,
-0.7455715092719436,
-0.7440696697400152,
-0.7426095484144338,
-0.7411900594073207,
-0.7398101410359031,
-0.7384687555121611,
-0.7371648886223161,
-0.7358975493972693,
-0.7346657697750326,
-0.7334686042561455,
-0.7323051295530186,
-0.7311744442340865,
-0.730075668363623,
-0.7290079431379773,
-0.7279704305190191,
-0.7269623128654421,
-0.7259827925626224,
-0.7250310916516166,
-0.7241064514578869,
-0.7232081322202865,
-0.722335412720795,
-0.7214875899154829,
-0.7206639785671193,
-0.7198639108798225,
-0.7190867361361372,
-0.7183318203368496,
-0.7175985458438696,
-0.716886311026466,
-0.7161945299110904,
-0.7155226318350579,
-0.7148700611042691,
-0.714236276655184,
-0.7136207517212114,
-0.7130229735036635,
-0.7124424428474267,
-0.7118786739214411,
-0.7113311939041225,
-0.7107995426737863,
-0.7102832725041672,
-0.7097819477650894,
-0.7092951446283301,
-0.7088224507787237,
-0.7083634651305372,
-0.707917797549109,
-0.707485068577799,
-0.7070649091702115,
-0.706656960427708,
-0.7062608733421848,
-0.705876308544093,
-0.7055029360556729,
-0.7051404350493755,
-0.7047884936114196,
-0.7044468085104555,
-0.7041150849712687,
-0.7037930364534946,
-0.7034803844352636,
-0.7031768582017391,
-0.702882194638474,
-0.7025961380295235,
-0.702318439860253,
-0.7020488586247651,
-0.7017871596378826,
-0.701533114851609,
-0.7012865026759999,
-0.7010471078043673,
-0.700814721042743,
-0.7005891391435222,
-0.7003701646432273,
-0.700157605704281,
-0.6999512759607593,
-0.699750994368002,
-0.6995565850560369,
-0.699367877186726,
-0.6991847048145584,
-0.6990069067510206,
-0.6988343264324628,
-0.6986668117913872,
-0.6985042151310924,
-0.6983463930035845,
-0.6981932060906986,
-0.698044519088352,
-0.69790020059385,
-0.6977601229961965,
-0.6976241623693097,
-0.6974921983681055,
-0.6973641141273568,
-0.6972397961632787,
-0.6971191342777676,
-0.6970020214652266,
-0.6968883538219259,
-0.6967780304578248,
-0.6966709534107974,
-0.6965670275632114,
-0.6964661605607833,
-0.6963682627336721,
-0.6962732470197457,
-0.6961810288899561,
-0.6960915262757942,
-0.6960046594987405,
-0.6959203512016872,
-0.695838526282262,
-0.6957591118280114,
-0.6956820370534018,
-0.6956072332385702,
-0.6955346336698072,
-0.6954641735816972,
-0.695395790100895,
-0.6953294221914829,
-0.6952650106018661,
-0.6952024978131742,
-0.6951418279891153,
-0.6950829469272576,
-0.6950258020116907,
-0.6949703421670095,
-0.6949165178137052,
-0.6948642808244588,
-0.6948135844830479,
-0.6947643834392965,
-0.694716633684565,
-0.6946702924556808,
-0.6946253184128031,
-0.6945816707944823,
-0.694539312416195,
-0.6944981982500809,
-0.6944583181200742,
-0.6944195363185361,
-0.694394640996012,
-0.6943821933347499],
'tck_k': 3,
'tck_t': [ 0.0,
0.0,
0.0,
0.0,
0.2,
0.4,
0.6,
0.8,
1.0,
1.2,
1.4,
1.6,
1.8,
2.0,
2.2,
2.4,
2.6,
2.8,
3.0,
3.2,
3.4,
3.6,
3.8,
4.0,
4.2,
4.4,
4.6,
4.8,
5.0,
5.2,
5.4,
5.6,
5.8,
6.0,
6.2,
6.4,
6.6,
6.8,
7.0,
7.2,
7.4,
7.6,
7.8,
8.0,
8.2,
8.4,
8.6,
8.8,
9.0,
9.2,
9.4,
9.6,
9.8,
10.0,
10.2,
10.4,
10.6,
10.8,
11.0,
11.2,
11.4,
11.6,
11.8,
12.0,
12.2,
12.4,
12.6,
12.8,
13.0,
13.2,
13.4,
13.6,
13.8,
14.0,
14.2,
14.4,
14.6,
14.8,
15.0,
15.2,
15.4,
15.6,
15.8,
16.0,
16.2,
16.4,
16.6,
16.8,
17.0,
17.2,
17.4,
17.6,
17.8,
18.0,
18.2,
18.4,
18.6,
18.8,
19.0,
19.2,
19.4,
19.6,
19.8,
20.0,
20.2,
20.4,
20.6,
20.8,
21.0,
21.2,
21.4,
21.6,
21.8,
22.0,
22.2,
22.4,
22.6,
22.8,
23.0,
23.2,
23.4,
23.6,
23.8,
24.0,
24.2,
24.4,
24.6,
24.8,
25.0,
25.2,
25.4,
25.6,
25.8,
26.0,
26.2,
26.4,
26.6,
26.8,
27.0,
27.2,
27.4,
27.6,
27.8,
28.0,
28.2,
28.4,
28.6,
28.8,
29.0,
29.2,
29.4,
29.6,
29.8,
30.0,
30.2,
30.4,
30.6,
30.8,
31.0,
31.2,
31.4,
31.6,
31.8,
32.0,
32.2,
32.4,
32.6,
32.8,
33.0,
33.2,
33.4,
33.6,
33.8,
34.0,
34.2,
34.4,
34.6,
34.8,
35.0,
35.2,
35.4,
35.6,
35.8,
36.0,
36.2,
36.4,
36.6,
36.8,
37.0,
37.2,
37.4,
37.6,
37.8,
38.0,
38.2,
38.4,
38.6,
38.8,
39.0,
39.2,
39.4,
39.6,
39.8,
40.0,
40.2,
40.4,
40.6,
40.8,
41.0,
41.2,
41.4,
41.6,
41.8,
42.0,
42.2,
42.4,
42.6,
42.8,
43.0,
43.2,
43.4,
43.6,
43.8,
44.0,
44.2,
44.4,
44.6,
44.8,
45.0,
45.2,
45.4,
45.6,
45.8,
46.0,
46.2,
46.4,
46.6,
46.8,
47.0,
47.2,
47.4,
47.6,
47.8,
48.0,
48.2,
48.4,
48.6,
48.8,
49.0,
49.2,
49.4,
49.6,
49.8,
50.0,
50.2,
50.4,
50.6,
50.8,
51.0,
51.2,
51.4,
51.6,
51.8,
52.0,
52.2,
52.4,
52.6,
52.8,
53.0,
53.2,
53.4,
53.6,
53.8,
54.0,
54.2,
54.4,
54.6,
54.8,
55.0,
55.2,
55.4,
55.6,
55.8,
56.0,
56.2,
56.4,
56.6,
56.8,
57.0,
57.2,
57.4,
57.6,
57.8,
58.0,
58.2,
58.4,
58.6,
58.8,
59.0,
59.2,
59.4,
59.6,
59.8,
60.0,
60.2,
60.4,
60.6,
60.8,
61.0,
61.2,
61.4,
61.6,
61.8,
62.0,
62.2,
62.4,
62.6,
62.8,
63.0,
63.2,
63.4,
63.6,
63.8,
64.0,
64.2,
64.4,
64.6,
64.8,
65.0,
65.2,
65.4,
65.6,
65.8,
66.0,
66.2,
66.4,
66.6,
66.8,
67.0,
67.2,
67.4,
67.6,
67.8,
68.0,
68.2,
68.4,
68.6,
68.8,
69.0,
69.2,
69.4,
69.6,
69.8,
70.0,
70.2,
70.4,
70.6,
70.8,
71.0,
71.2,
71.4,
71.6,
71.8,
72.0,
72.2,
72.4,
72.6,
72.8,
73.0,
73.2,
73.4,
73.6,
73.8,
74.0,
74.2,
74.4,
74.6,
74.8,
75.0,
75.2,
75.4,
75.6,
75.8,
76.0,
76.2,
76.4,
76.6,
76.8,
77.0,
77.2,
77.4,
77.6,
77.8,
78.0,
78.2,
78.4,
78.6,
78.8,
79.0,
79.2,
79.4,
79.6,
79.8,
80.0,
80.2,
80.4,
80.6,
80.8,
81.0,
81.2,
81.4,
81.6,
81.8,
82.0,
82.2,
82.4,
82.6,
82.8,
83.0,
83.2,
83.4,
83.6,
83.8,
84.0,
84.2,
84.4,
84.6,
84.8,
85.0,
85.2,
85.4,
85.6,
85.8,
86.0,
86.2,
86.4,
86.6,
86.8,
87.0,
87.2,
87.4,
87.6,
87.8,
87.8,
87.8,
87.8],
'time_at_umax': 36.17889447236181}}}
Extract Growth Statistics#
# Extract stats from each non-parametric fit
stats_spline = gc.inference.extract_stats(
fit_spline,
t,
N,
phase_boundary_method="tangent",
)
stats_sliding_window = gc.inference.extract_stats(
fit_sliding_window,
t,
N,
phase_boundary_method="tangent",
)
# Combine stats into a dictionary
phenom_nonparam_stats = {
"spline": stats_spline,
"sliding_window": stats_sliding_window,
}
# Create comparison dataframe
print("=== Phenomenological Non-Parametric Models Comparison ===")
phenom_nonparam_df = pd.DataFrame(phenom_nonparam_stats).T[
[
"mu_max",
"intrinsic_growth_rate",
"doubling_time",
"time_at_umax",
"exp_phase_start",
"exp_phase_end",
"model_rmse",
]
]
phenom_nonparam_df
=== Phenomenological Non-Parametric Models Comparison ===
| mu_max | intrinsic_growth_rate | doubling_time | time_at_umax | exp_phase_start | exp_phase_end | model_rmse | |
|---|---|---|---|---|---|---|---|
| spline | 0.077924 | None | 8.895163 | 36.178894 | 21.572282 | 50.946358 | 0.0 |
| sliding_window | 0.077894 | None | 8.898575 | 36.2 | 21.56668 | 50.952021 | 0.000005 |
Customizing Phase Boundary Detection#
Two methods are available for determining exponential phase boundaries:
1. Threshold Method#
Tracks the instantaneous specific growth rate μ(t)
exp_phase_start: First time when μ exceeds a fraction of μ_max (default: 15%)exp_phase_end: First time after peak when μ drops below the threshold
2. Tangent Method#
Constructs a tangent line in log space at the point of maximum growth rate
Extends this tangent to intersect baseline (exp_phase_start) and plateau (exp_phase_end)
# Compare phase-boundary methods on the same fit
phase_boundary_rows = []
# Tangent method
stats_tangent = gc.inference.extract_stats(
fit_spline,
t,
N,
phase_boundary_method="tangent",
)
phase_boundary_rows.append(
{
"label": "tangent",
"method": "tangent",
"lag_threshold": np.nan,
"exp_threshold": np.nan,
"stats": stats_tangent,
}
)
# Threshold method at different cutoffs
for frac, label in [(0.10, "threshold_low"), (0.30, "threshold_high")]:
stats_threshold = gc.inference.extract_stats(
fit_spline,
t,
N,
phase_boundary_method="threshold",
lag_threshold=frac,
exp_threshold=frac,
)
phase_boundary_rows.append(
{
"label": label,
"method": "threshold",
"lag_threshold": frac,
"exp_threshold": frac,
"stats": stats_threshold,
}
)
# Create comparison dataframe
print("=== Phase Boundary Method Comparison ===")
phase_boundary_df = pd.DataFrame(
[
{
"label": row["label"],
"method": row["method"],
"lag_threshold": row["lag_threshold"],
"exp_threshold": row["exp_threshold"],
"exp_phase_start": row["stats"]["exp_phase_start"],
"exp_phase_end": row["stats"]["exp_phase_end"],
}
for row in phase_boundary_rows
]
)
phase_boundary_df
=== Phase Boundary Method Comparison ===
| label | method | lag_threshold | exp_threshold | exp_phase_start | exp_phase_end | |
|---|---|---|---|---|---|---|
| 0 | tangent | tangent | NaN | NaN | 21.572282 | 50.946358 |
| 1 | threshold_low | threshold | 0.1 | 0.1 | 9.877993 | 62.484526 |
| 2 | threshold_high | threshold | 0.3 | 0.3 | 18.206304 | 54.130940 |
See more details on the phase boundary methods in the next tutorial notebook:
plotting.ipynb (Visualize fitted growth curves, derivatives,
and growth statistics)
